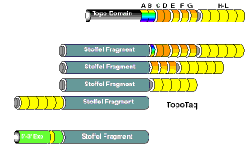
TopoTaq
 |
Contact info: Fidelity Systems 7961 Cessna Avenue Gaithersburg MD 20879-4117 301-527-0804 (tel) 301-527-8250 (fax) fsi1@fidelitysystems.com |

The Complete Genome
of the Hyperthermophile
Methanopyrus kandleri AV19
and Monophyly of
Archaeal Methanogens
Slesarev,A.I., Mezhevaya,K.V., Makarova,K.S.,
Polushin,N.N., Shcherbinina,O.V.,
Shakhova,V.V., Belova,G.I., Aravind,L.,
Natale,D.A., Rogozin,I.B., Tatusov,R.L.,
Wolf,Y.I., Stetter,K.O., Malykh,A.G.,
Koonin,E.V. and Kozyavkin,S.A. (2002)
Proc. Natl. Acad. Sci. U.S.A. 99, 4644-4649
EVOLUTION: Stripped Down to the Bare Essentials
Science 296, 431 (2002) Editors' Choice
A HAPPY MARRIAGE: advancing DNA polymerases with DNA topoisomerase
Trends in Biotech 20, 491 (2002) Journal Club
Direct Automated Sequencing of Bacterial Genomic DNA
At the Ninth International Genome Sequencing and Analysis Conference Dr. Bruce Roe from the University of Oklahoma presented his group's data on the successful automated sequencing of non-clonable and non-PCRable regions of Streptococcus pyogenes (1.92 Mb). The scientists used ThermoFidelase I and a new sequencing kit to sequence genomic DNA directly. This took less than a week from receiving new reagents to presenting the most stunning results to 1200 attendees of the conference. "It completely changed our finishing strategy" - said Dr. Roe.
Successful Sequencing of Non-sequenceable DNA
At the 10th Annual Cold Spring Harbor Meeting on Genome Mapping and Sequencing Dr. Elaine Mardis of Washington University presented data on using ThermoFidelase I in large scale sequencing projects. "This approach has proven successful in sequencing through stop regions of M13 or pUC subclones, and PCR products" - concluded Dr. Mardis.
(c) 1996-2014 Fidelity Systems