Fluorescence Detection of Sequence-Specific Annealing of DNA Probes
ORDERING INFO
(2004) NAR 32 e62 (2001) CBB 34 257 (review) (1998) SPIE 3256 68 (1997) Anal. Biochem 244 86 (1997) Biophys J 73 3277 (1995) NAR 23 2872 CONTACT INFO 301-527-0804 (tel) 301-527-8250 (fax) fsi1@fidelitysystems.com |
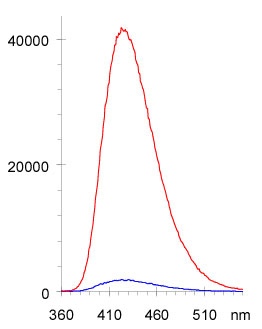
Most of the quenching (loss of fluorescence signal) of the nucleoside analogs is associated with incorporation into a single stranded oligonucleotide. Since this phenomenon is apparently associated with base stacking interactions, we investigated the fluorescence of these probes after deliberate disruption of base stacking through the formation of a bulge or hairpin. In this application the probe is positioned as an insertion in a sequence that is complementary to a sequence of interest. Since the fluorophore does not have a base-pairing partner in the target sequence, when annealing occurs, it is squeezed out of the strand to form a single base hairpin and this results in a strong increase in fluorescence intensity. The fluorescence change associated with annealing of the two strands is dependent on the identity of bases neighboring the probe and produces an increase of up to 27 times the background signal (equimolar single strand containing the probe). The associated depression of melting temperature for the resulting hairpin or bulge in a 21-mer is 1 deg C, an indication that there is little perturbation in annealing caused by the probe. Figure shows the type of increase in intensity observed in this hairpin hybridization application. This approach allows one to probe for a specific sequence within a mixture without separation of products.
These probes can be incorporated into oligonucleotides site-specifically and since they are more heavily quenched by neighboring purines, and much less quenched by neighboring pyrimidines, they can be selectively placed within an oligonucleotide to get a brighter or quieter response. Oligonucleotides containing the probe in a terminal position or containing multiple probes per strand are extremely bright. A 21 fold increase in fluorescence: 3-MI in a bulge of the double stranded oligo vs 3-MI in the single stranded oligo.
FOR RESEARCH USE ONLY |